CLARK
Taxonomic ClassificationCLARK (CLAssifier based on Reduced K-mers) is a supervised sequence classification method that uses discriminative k-mers — k-mers unique to a specific taxon — for fast and precise metagenomic classification. [1]
How to Obtain Output Model File
Below is a brief workflow the team ran to obtain the output model examples we present on the tools page.
Input
FASTQ/FASTA files (single-end or paired-end reads)
Output
CSV classification results with confidence scores and estimated abundance table
conda install -c bioconda clark
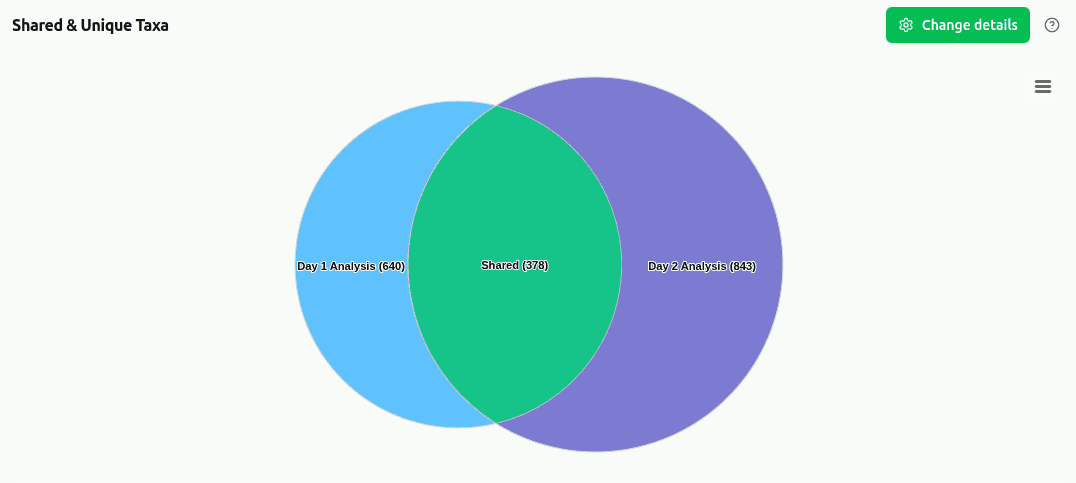
Sample 1 human gut metagenome (SRR14092160, 5M read subset) classified with CLARK-l against a custom Database of 31 gut-associated bacterial genomes.
- 1
Download reads from NCBI SRA
prefetch SRR14092160 && fasterq-dump SRR14092160 -O /data --split-files && gzip /data/SRR14092160_*.fastq
Human gut metagenome, pre-VRE colonization timepoint (Day -9). Illumina paired-end, ~5.5M read pairs (partial extraction).
- 2
Subset to 5M reads and decompress
zcat SRR14092160_1.fastq.gz | head -20000000 > subset_1.fastq && zcat SRR14092160_2.fastq.gz | head -20000000 > subset_2.fastq
CLARK does not support gzipped FASTQ. 5M reads = 20M lines in FASTQ format.
- 3
Build custom 31-genome database
Download 31 RefSeq genomes via NCBI Datasets, prepare targets.txt and NCBI taxonomy files (nodes.dmp, names.dmp).
- 4
Run CLARK-l classification
CLARK-l -T /db/targets.txt -D /db/custom_0/ -P subset_1.fastq subset_2.fastq -R clark_result -n 8 -m 0
CLARK-l is the light-weight mode (~4 GB RAM). 0.20% classified (10,049 / 5,000,000 reads). 26 species detected.
- 5
Estimate abundance
estimate_abundance.sh -F clark_result.csv -D /db --highconfidence
Produces the CSV abundance file that IntMeta reads. The --highconfidence flag filters low-confidence assignments.
clark_abundance.clark to IntMetaMaterials Used
Charts Reference
Detailed descriptions for all 12 visualizations generated by CLARK in IntMeta.
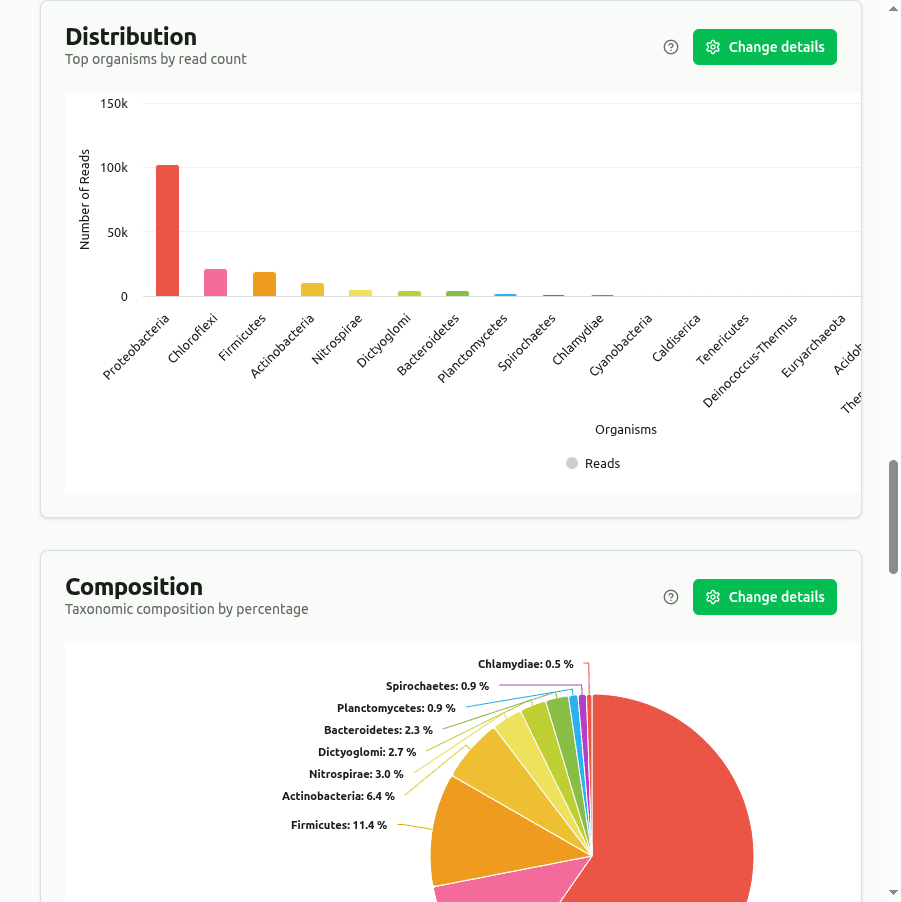
distribution
Bar chart of the top organisms ranked by read count from CLARK's discriminative k-mer classification. Only reads matching k-mers unique to a single taxon are counted, producing high-confidence assignments.
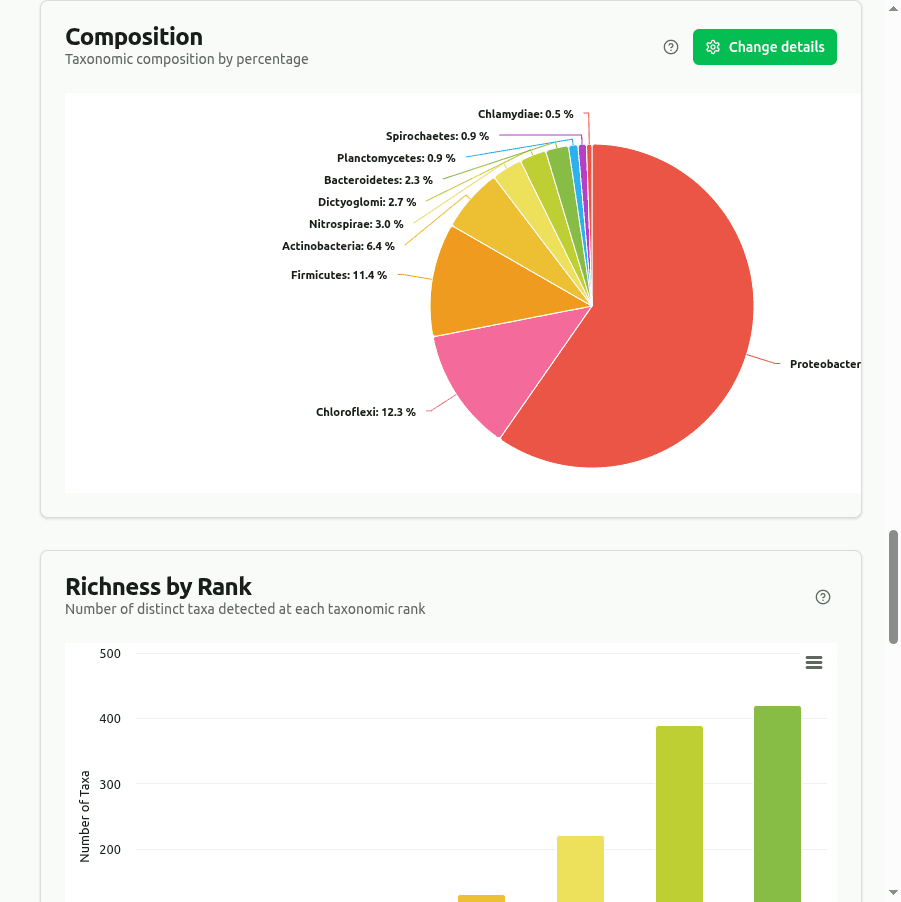
composition
Pie/donut chart showing the top classified taxa as a percentage of their combined read count. Each slice is (taxon reads / sum of displayed reads) × 100 at the selected rank.
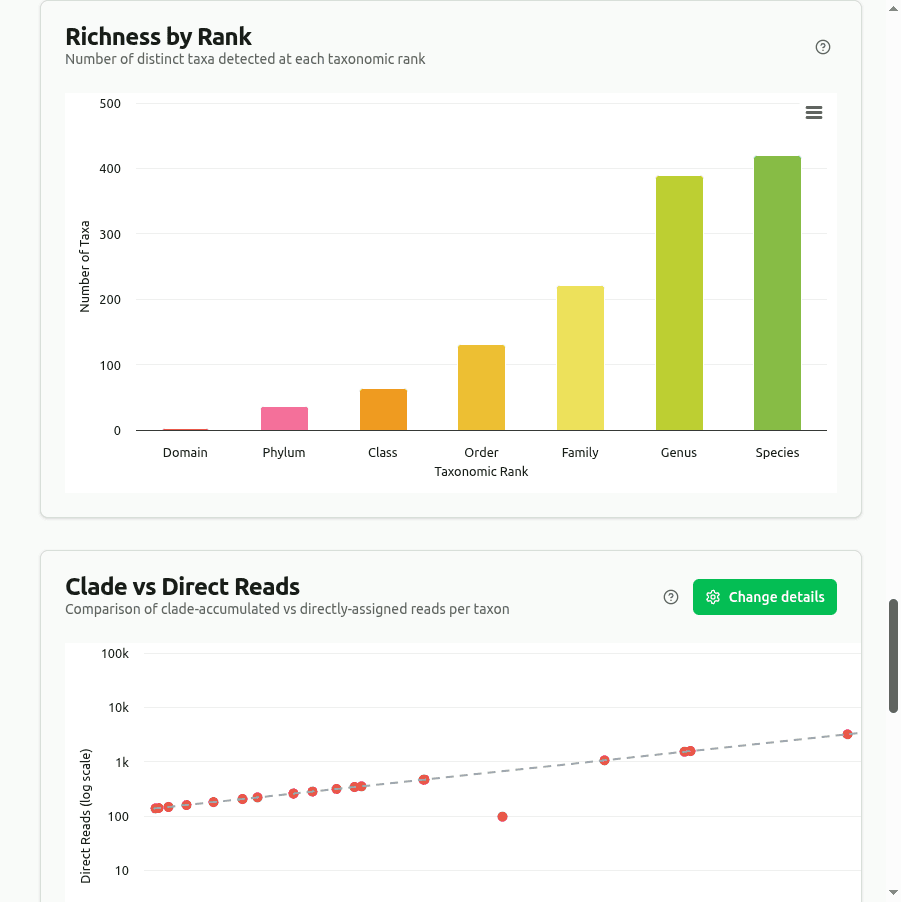
richness
Counts every distinct organism with at least one assigned read at each major rank (Domain through Species). CLARK's discriminative k-mer approach typically reports fewer taxa than Kraken2, but each assignment carries higher specificity.
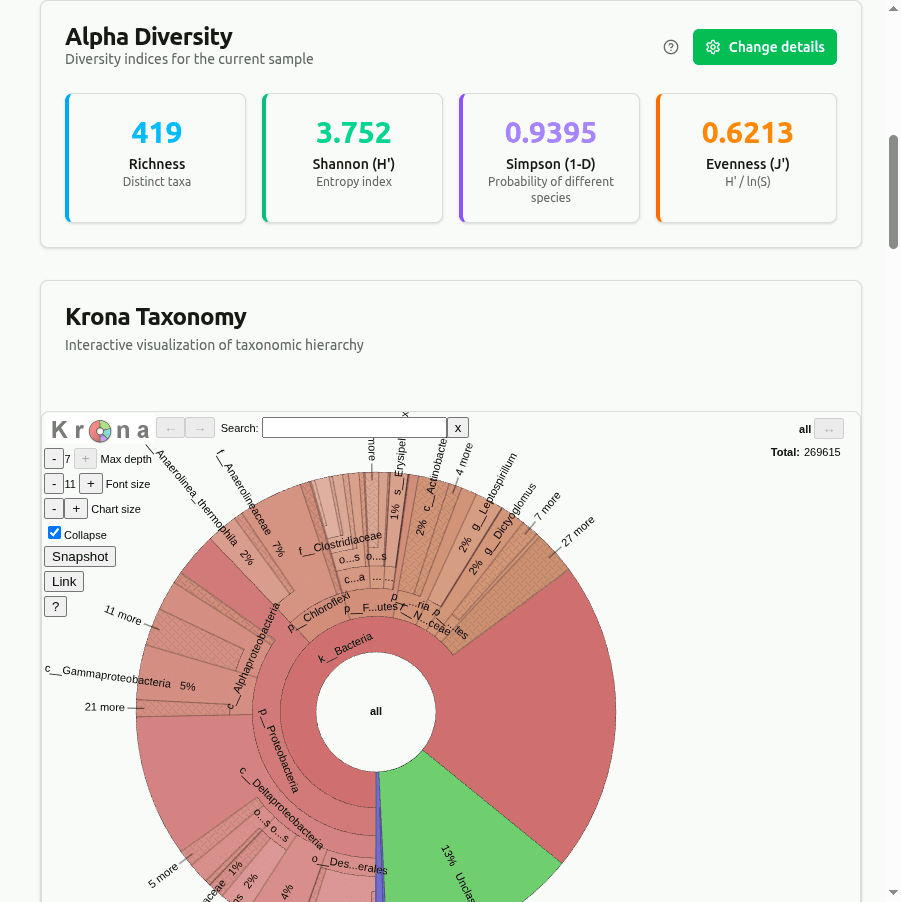
diversity
Alpha diversity indices computed from per-taxon read-count proportions: Shannon H = −Σ(pᵢ · ln pᵢ), Simpson D = 1 − Σ(nᵢ(nᵢ−1))/(N(N−1)), and Pielou's evenness J = H / ln(richness). Useful for comparing community evenness across samples.
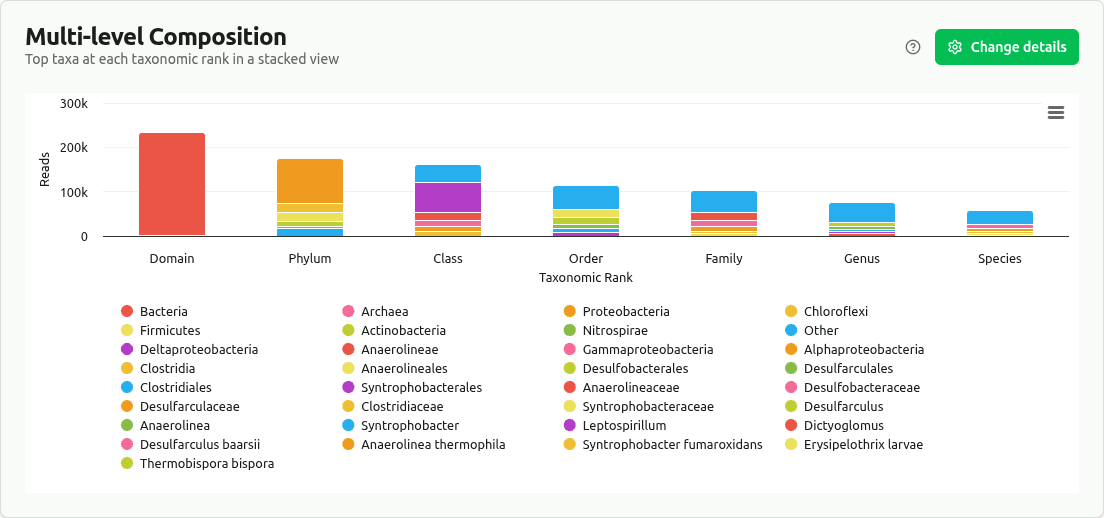
multilevel-composition
Stacked bar chart with the top taxa (by read count) at each major rank; remaining taxa grouped as 'Other'. Shows how the dominant organisms shift across classification depths.
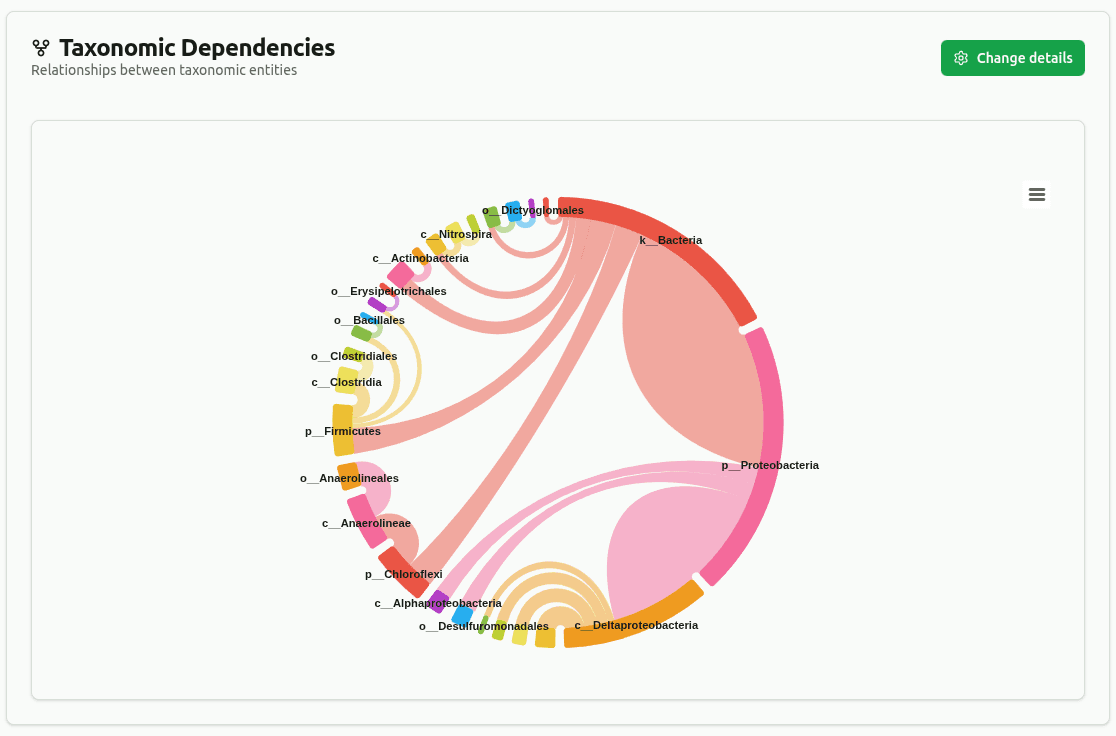
dependency-wheel
Chord diagram linking parent to child taxa across ranks, with connection thickness proportional to shared read count. Edges below the minimum coverage threshold (default 5%) are filtered out.
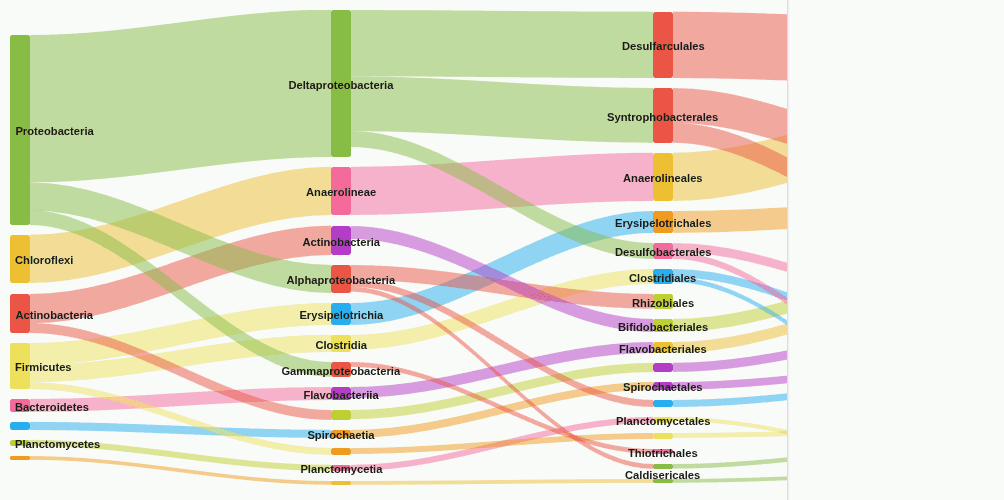
sankey-flow
Sankey flow diagram tracing how reads distribute from the start rank to the end rank through the lineage hierarchy. Band width equals read count between connected taxa; low-coverage edges are removed.
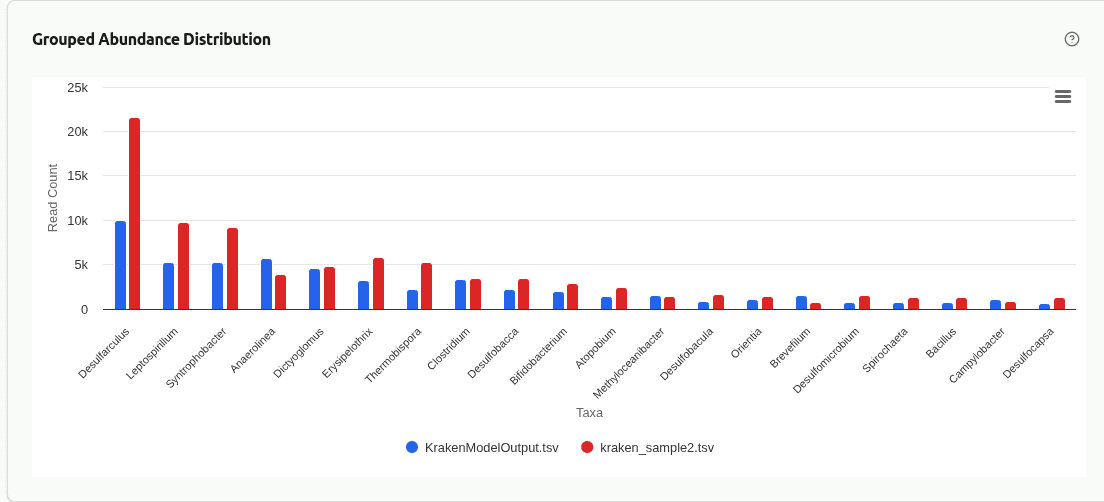
comp-grouped-abundance
Grouped bar chart placing the top taxa from each sample side by side at the selected rank. Each group contains one bar per sample, colored by sample identity, enabling direct visual comparison of absolute read counts for the same organism across samples.
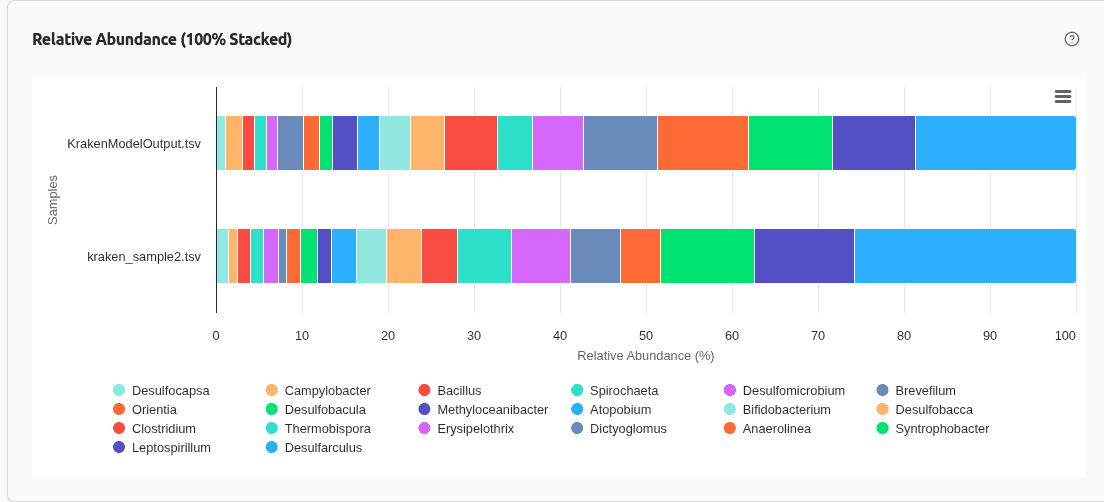
comp-relative-abundance
100% stacked bar chart where each bar represents one sample and segments show the proportional contribution of each taxon. Useful for comparing community composition when samples have very different sequencing depths, since all bars are normalized to the same height.

comp-abundance-heatmap
Color-matrix heatmap with taxa on one axis and samples on the other. Cell color intensity is proportional to read abundance at the selected rank. Hierarchical clustering on both axes groups similar samples and co-occurring taxa together.
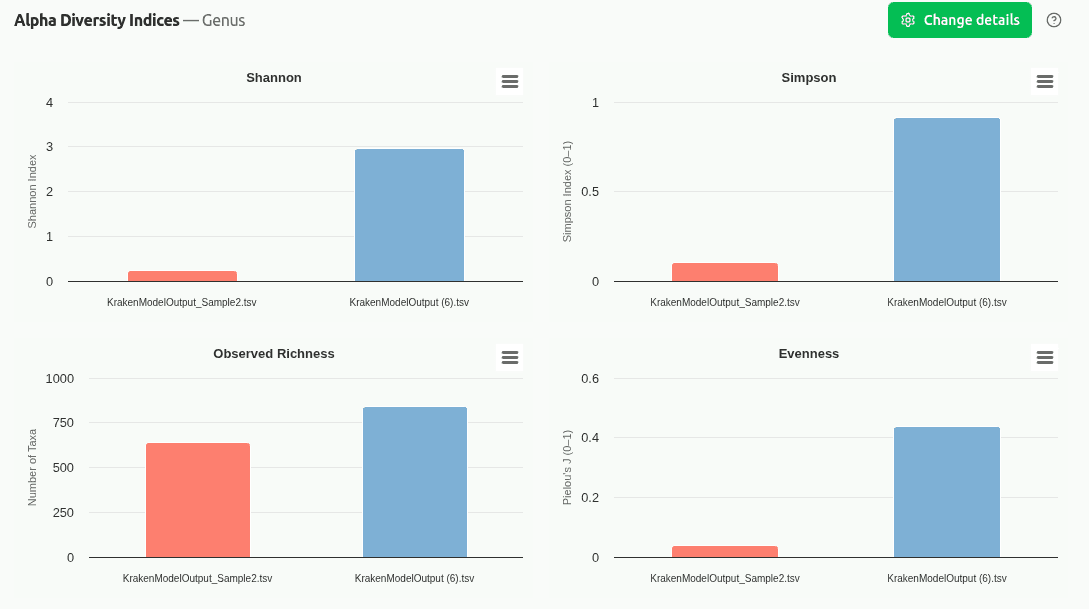
comp-diversity-indices
Multi-panel chart displaying Shannon entropy, Simpson diversity, Observed Richness, and Pielou's Evenness for each sample on its own y-axis scale. Enables quick cross-sample comparison of alpha diversity without scale distortion.